Introduction
Classification tree methods (i.e., decision tree methods) are recommended when the data mining task contains classifications or predictions of outcomes, and the goal is to generate rules that can be easily explained and translated into SQL or a natural query language.
A Classification tree labels, records, and assigns variables to discrete classes. A Classification tree can also provide a measure of confidence that the classification is correct.
A Classification tree is built through a process known as binary recursive partitioning. This is an iterative process of splitting the data into partitions, and then splitting it up further on each of the branches.
Initially, a Training Set is created where the classification label (i.e., purchaser or non-purchaser) is known (pre-classified) for each record. Next, the algorithm systematically assigns each record to one of two subsets on the some basis (i.e., income > $75,000 or income <= $75,000). The object is to attain an homogeneous set of labels (i.e., purchaser or non-purchaser) in each partition. This partitioning (splitting) is then applied to each of the new partitions. The process continues until no more useful splits can be found. The heart of the algorithm is the rule that determines the initial split rule (displayed in the following figure).
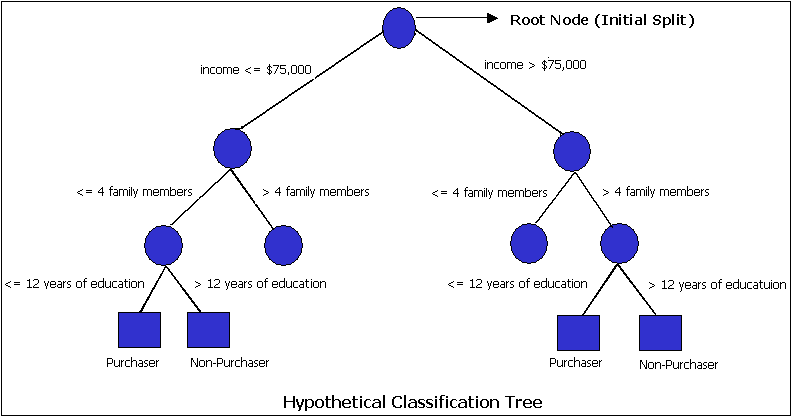
The process starts with a Training Set consisting of pre-classified records (target field or dependent variable with a known class or label such as purchaser or non-purchaser). The goal is to build a tree that distinguishes among the classes. For simplicity, assume that there are only two target classes, and that each split is a binary partition. The partition (splitting) criterion generalizes to multiple classes, and any multi-way partitioning can be achieved through repeated binary splits. To choose the best splitter at a node, the algorithm considers each input field in turn. In essence, each field is sorted. Every possible split is tried and considered, and the best split is the one that produces the largest decrease in diversity of the classification label within each partition (i.e., the increase in homogeneity). This is repeated for all fields, and the winner is chosen as the best splitter for that node. The process is continued at subsequent nodes until a full tree is generated.
Analytic Solver Data Science uses the Gini index as the splitting criterion, which is a commonly used measure of inequality. The index fluctuates between a value of 0 and 1. A Gini index of 0 indicates that all records in the node belong to the same category. A Gini index of 1 indicates that each record in the node belongs to a different category. For a complete discussion of this index, please see Leo Breiman’s and Richard Friedman’s book, Classification and Regression Trees (3).
Pruning the Tree
Pruning is the process of removing leaves and branches to improve the performance of the decision tree when moving from the Training Set (where the classification is known) to real-world applications (where the classification is unknown). The tree-building algorithm makes the best split at the root node where there are the largest number of records, and considerable information. Each subsequent split has a smaller and less representative population with which to work. Towards the end, idiosyncrasies of training records at a particular node display patterns that are peculiar only to those records. These patterns can become meaningless for prediction if you try to extend rules based on them to larger populations.
For example, if the classification tree is trying to predict height and it comes to a node containing one tall person X and several other shorter people, the algorithm decreases diversity at that node by a new rule imposing people named X are tall, and thus classify the Training Data. In the real world, this rule is obviously inappropriate. Pruning methods solve this problem -- they let the tree grow to maximum size, then remove smaller branches that fail to generalize. (Note: Do not include irrelevant fields such as name, as this is simply used an illustration.)
Since the tree is grown from the Training Set, when it has reaches full structure it usually suffers from over-fitting (i.e., it is explaining random elements of the Training Data that are not likely to be features of the larger population of data). This results in poor performance on data. Therefore, trees must be pruned using the Validation Set.
Ensemble Methods
Analytic Solver Data Science offers three powerful ensemble methods for use with Classification trees: bagging (bootstrap aggregating), boosting, and random trees. The Classification Tree Algorithm on its own can be used to find one model that results in good classifications of the new data. We can view the statistics and confusion matrices of the current classifier to see if our model is a good fit to the data, but how would we know if there is a better classifier waiting to be found? The answer is that we do not know if a better classifier exists. However, ensemble methods allow us to combine multiple weak classification tree models that, when taken together form a new, more accurate strong classification tree model. These methods work by creating multiple diverse classification models, taking different samples of the original data set, and then combining their outputs. Outputs may be combined by several techniques for example, majority vote for classification and averaging for regression. This combination of models effectively reduces the variance in the strong model. The three different type of ensemble methods offered in Analytic Solver Data Science (bagging, boosting, and random trees) differ on three items: 1) the selection of a Training Set for each classifier or weak model; 2) how the weak models are generated; and 3) how the outputs are combined. In all three methods, each weak model is trained on the entire Training Set to become proficient in some portion of the data set.
Bagging (bootstrap aggregating) was one of the first ensemble algorithms to be documented. It is a simple, effective algorithm. Bagging generates several Training Sets by using random sampling with replacement (bootstrap sampling), applies the classification tree algorithm to each data set, then takes the majority vote between the models to determine the classification of the new data. The biggest advantage of bagging is the relative ease with which the algorithm can be parallelized, which makes it a better selection for very large data sets.
Boosting builds a strong model by successively training models to concentrate on the misclassified records in previous models. Once completed, all classifiers are combined by a weighted majority vote. Analytic Solver Data Science offers three different variations of boosting as implemented by the AdaBoost algorithm (ensemble algorithm): M1 (Freund), M1 (Breiman), and SAMME (Stagewise Additive Modeling using a Multi-class Exponential).
Adaboost.M1 first assigns a weight (wb(i)) to each record or observation. This weight is originally set to 1/n, and is updated on each iteration of the algorithm. An original classification tree is created using this first Training Set (Tb) and an error is calculated as
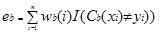
where, the I() function returns 1 if true, and 0 if not.
The error of the classification tree in the bth iteration is used to calculate the constant ?b. This constant is used to update the weight (wb(i). In AdaBoost.M1 (Freund), the constant is calculated as
αb= ln((1-eb)/eb)
In AdaBoost.M1 (Breiman), the constant is calculated as
αb= 1/2ln((1-eb)/eb)
In SAMME, the constant is calculated as
αb= 1/2ln((1-eb)/eb + ln(k-1) where k is the number of classes
where, the number of categories is equal to 2, SAMME behaves the same as AdaBoost Breiman.
In any of the three implementations (Freund, Breiman, or SAMME), the new weight for the (b + 1)th iteration will be
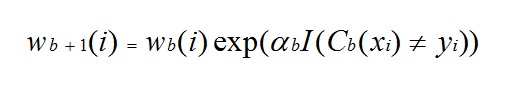
Afterwards, the weights are all readjusted to sum to 1. As a result, the weights assigned to the observations that were classified incorrectly are increased and the weights assigned to the observations that were classified correctly are decreased. This adjustment forces the next classification model to put more emphasis on the records that were misclassified. (The ? constant is also used in the final calculation, which will give the classification model with the lowest error more influence.) This process repeats until b = Number of weak learners (controlled by the User). The algorithm then computes the weighted sum of votes for each class and assigns the winning classification to the record. Boosting generally yields better models than bagging; however, it does have a disadvantage as it is not parallelizable. As a result, if the number of weak learners is large, boosting would not be suitable.
Random trees (i.e., random forests) is a variation of bagging. This method works by training multiple weak classification trees using a fixed number of randomly selected features (sqrt[number of features] for classification, and a number of features/3 for prediction), then takes the mode of each class to create a strong classifier. Typically, in this method the number of “weak” trees generated could range from several hundred to several thousand depending on the size and difficulty of the training set. Random Trees are parallelizable since they are a variant of bagging. However, since Random Trees selects a limited amount of features in each iteration, the performance of random trees is faster than bagging.
Classification Tree Ensemble methods are very powerful methods, and typically result in better performance than a single tree. This feature addition provides more accurate classification models and should be considered over the single tree method.
